Using language models to design antibodies to combat autoimmune disorders
TL;DR: We adapted our protein language model ProGen to optimize antibodies that bind to a protein called “CD40L”, a critical target for autoimmune disorders. We tested our AI designed antibodies in the laboratory and found that they bound very tightly to CD40L, showcasing the potential of this approach for antibody drug development.
Salesforce AI Research has been adopting its advances in NLP for applications in protein design under our AI for Social Good charter. We developed the protein language model ProGen with the hope that it would be a step towards using AI to develop treatments for diseases. In our previous work, we trained ProGen on antibacterial lysozyme proteins, and found that it could design very novel lysozymes that retained antibacterial properties. In this work, we go a step beyond that and use ProGen to help design antibody proteins with direct clinical applications. Please refer to our paper for full details of this work.
Antibodies play a pivotal role in our body's defense against infections. These tiny molecules are crafted by our immune system to latch onto foreign invaders, known as antigens. This process is crucial for our immune system's proper functioning. Laboratory-developed antibodies have proven to be highly effective in treating various diseases, with more than 100 antibody treatments approved by the FDA. In this work, we used AI to design nanobodies, which are a special class of antibodies that are composed of a single protein.
The target system for the antibodies designed in this study involves 2 proteins: CD40 and CD40L. These proteins play a critical role in regulating the immune system. They work by bringing together T-cells and antigen-presenting cells to trigger an immune response.
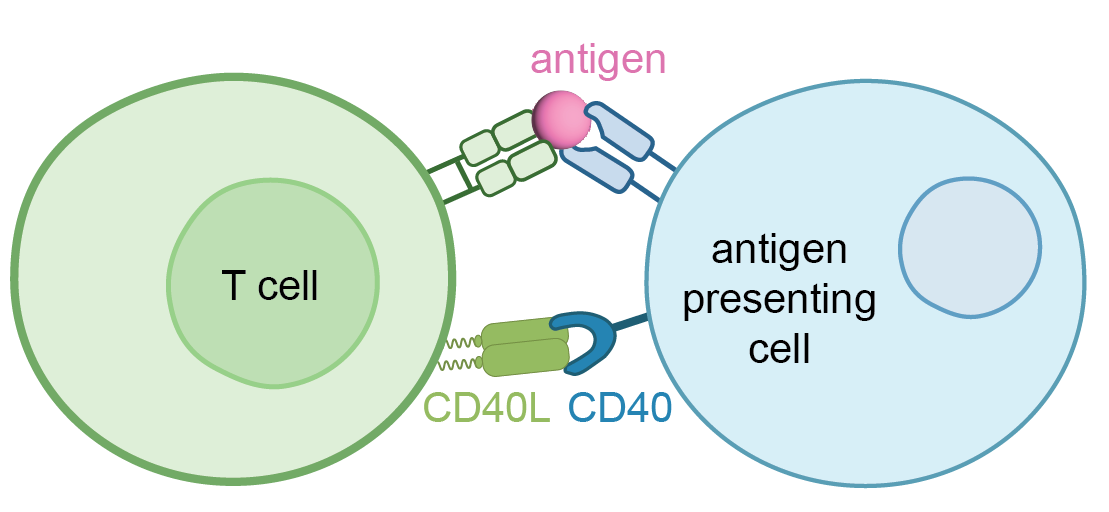
In individuals with autoimmune disorders, this immune response can become out of control, causing the body to attack itself. To address this issue, we aimed to disrupt the interaction between CD40 and CD40L. One way to do this is by developing an antibody that binds strongly to CD40L, making it harder for CD40 to connect with CD40L. Our objective was to employ AI to identify antibodies that exhibit high binding affinity to CD40L.
Our collaborators at Twist Bioscience collected data by screening a large set of random antibodies for their binding capacity to CD40L. This initial data provided us with a valuable starting point for designing new antibodies.
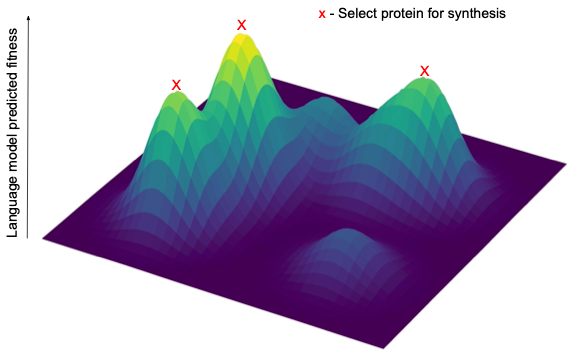
We employed various techniques to train AI language models to predict the binding affinity of new antibodies to CD40L. These techniques included training our in-house protein language model, ProGen, to understand the characteristics of the most effective antibodies in the lab data. We also used other methods to predict binding affinity from the feature representations learned by protein language models. Combining all of these models, we created a scoring system to map out the fitness landscape of these antibodies, helping us select a diverse set of high-potential antibodies for further testing.
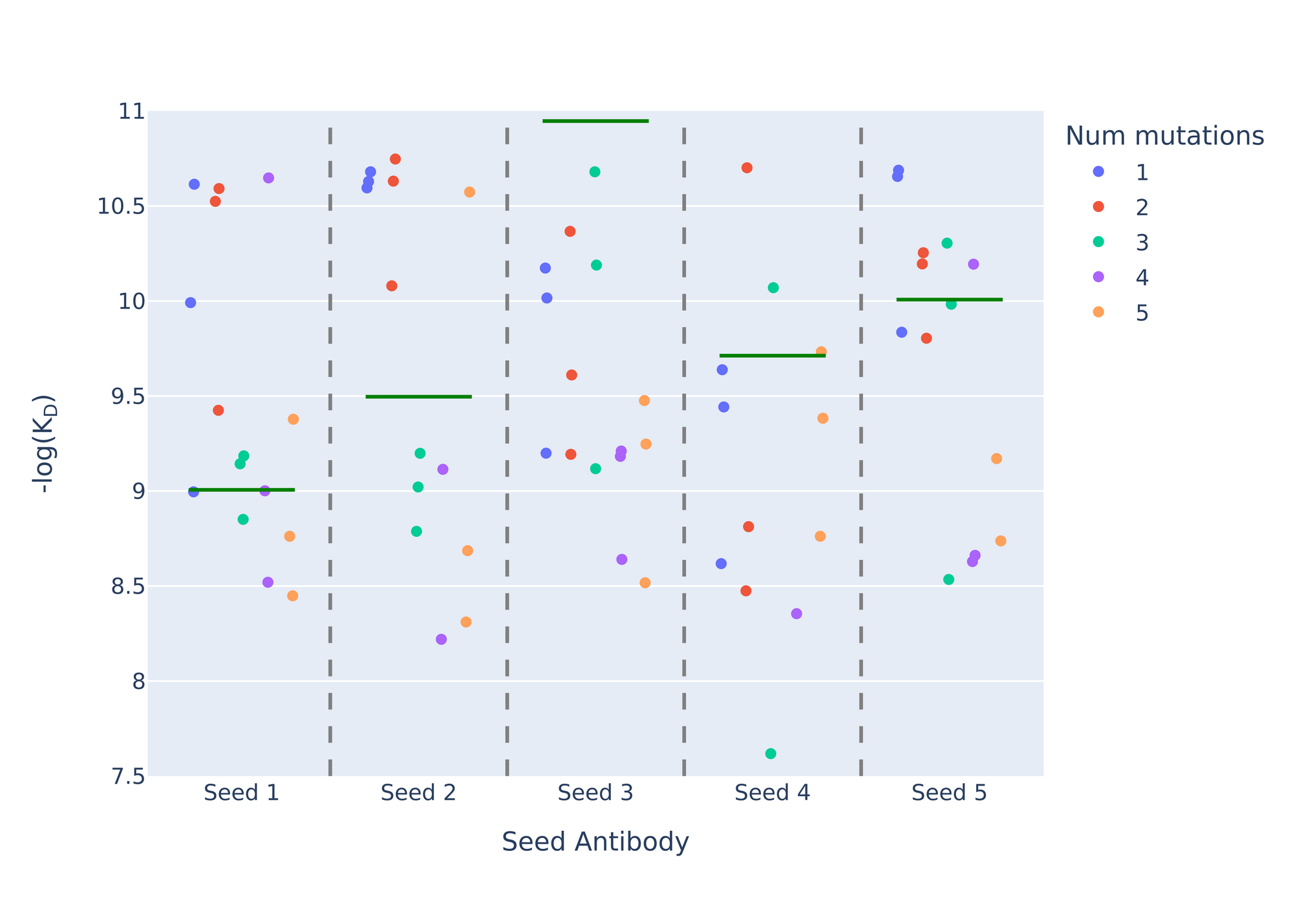
To begin, we took five antibodies from the initial lab experiments, which bound tightly to CD40L, and treated them as "seed antibodies" for the AI to modify. The strongest binder among these five starting antibodies had an affinity at the detection limit, indicating its binding strength exceeded the measurement capabilities of the lab instruments. The weakest seed antibody was about 40-50 times weaker than the limit of detection, which is still fairly strong. The remaining three seed antibodies fell within this range. Using our AI models, we designed 15 optimized antibodies for each of these five seed antibodies, totaling 75 new antibody designs. We succeeded in optimizing all five seed antibodies to reach the detection limit, regardless of their initial binding strength.
In addition to these, we designed 21 other antibodies with more flexibility, not tied to a specific seed antibody. Instead, they combined different antibody sequence spans known as Complementarity-Determining Regions, or "CDRs", in novel ways. Within this group, we discovered a couple more antibodies that reached the detection limit, as well as strong binders that were up to 8 mutations away from the closest antibody in our training set.
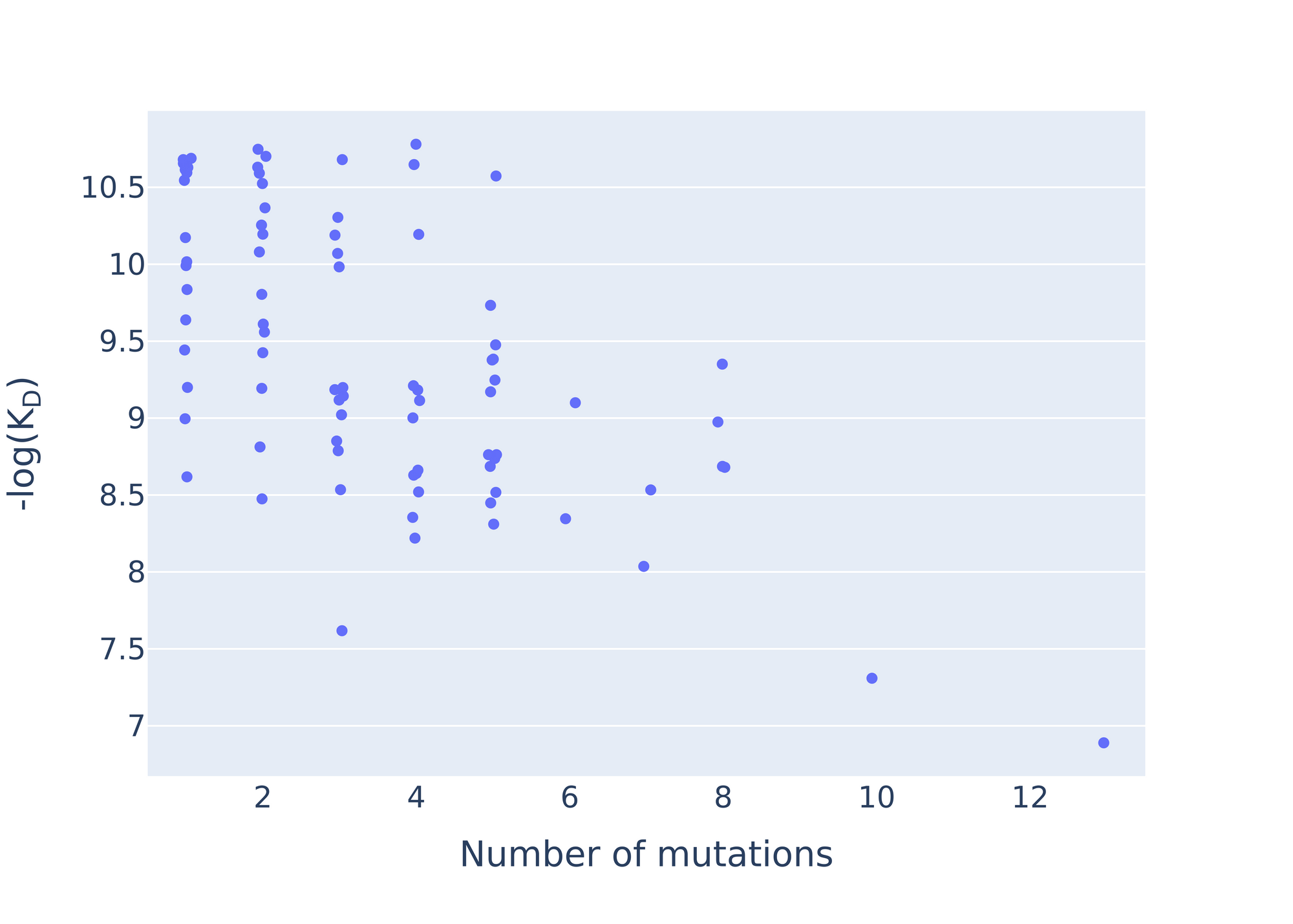
Our methods were able to optimize five antibodies to reach the detection limit, regardless of their starting affinity, and discover numerous potent CD40L binders that hold promise as potential treatments for autoimmune disorders. Our approach could also be applied to help develop antibodies that bind to other targets and treat other diseases. We are excited about the possibilities that this work unlocks, as it brings the field closer to using AI language models to develop therapeutics for diseases.
Authors and acknowledgements
This was a joint collaboration between Salesforce AI Research, Twist Bioscience, and Vanderbilt University School of Medicine, with co-authors Ben Krause, Subu Subramanian, Tom Yuan, Marisa Yang, Aaron Sato, and Nikhil Naik. We also thank Ali Madani, Richard Socher, Vanita Nemali, Audrey Cook, Caiming Xiong, and Silvio Saverese, Hoa Giang, Maxwell Stefan, Joyce Lai, and Sejal Petal for their support.